The DNA in our cells constantly
undergoes different types of damage, but fortunately mechanisms have evolved
that can repair this damage. However, if DNA damage is not repaired this can
lead to mutations, cancer or cell death. In recent years, research
investigating different mechanisms of DNA repair has led to a better
understanding of this process. My research studies a unique organism that has
an unprecedented DNA repair system among animals. These small animals
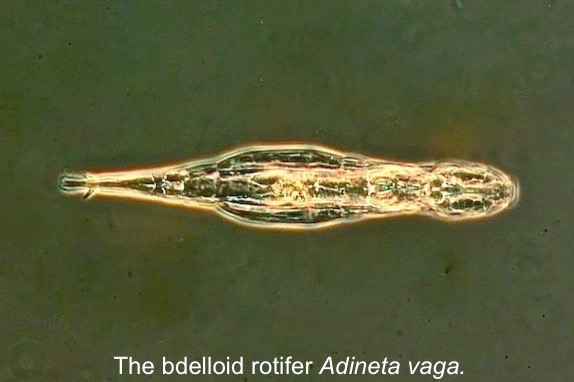
(bdelloids rotifers) repair DNA damage caused by doses of ionizing radiation
that are >100x greater than the lethal dose for humans. These rotifers are
therefore a unique model for studying DNA repair.
The objective of my research is to
identify genes, proteins and regulatory mechanisms involved in DNA repair in
bdelloids rotifers. Current projects in my lab involve:
- Developing the tools for
using CRISPR/Cas9 genome editing to knock out candidate DNA repair genes in
bdelloids.
- Using RNA interference
(RNAi) to knock down expression of target genes. This is done by engineering
strains of bacteria that are then fed to rotifers to induce RNAi and silence
gene expression.
- Using yeast two-hybrid
analysis to characterize protein interactions among DNA repair rotifers in
rotifers.
-
Investigating the evolution
of histone genes in rotifer genomes and the roles of novel histone variants and
epigenetics in the regulation of DNA repair and other processes.
Current students in the lab:
Chelbi Gilmore (BCMB '20)
Connor Onitsuka (BCMB '20)
Storm Gardner (BCMB '19)
Tristian Wiles (BCMB '21)
Rose Johnson (Biology '21)
Mitchell Rotenberry (BCMB '21)
Publications
- Schurko,
A.M. 2013. To “bee or not to bee” male of female? An educational primer for use with “The Am-tra1 gene is an essential regulator of female splice regulation at two levels of the sex determination hierarchy of the honeybee.” Genetics (In Press).
- Hanson, S., Schurko, A.M., Hecox-Lea, B., Mark Welch, D., Stelzer, C.-P. and Logsdon, J. Inventory and phylogenetic
analysis of meiotic genes in monogonont rotifers. Journal of Heredity(Accepted Feb. 11, 2013).
- Schurko, A.M., Mazur, D.J. and Logsdon, Jr., J.M. 2010. Inventory and phylogenomic distribution of meiotic genes in Nasonia vitripennis and among diverse arthropods. Insect Molecular Biology 19:165-80.
- Werren, J.H., Richards, S., Desjardins, C.A., Niehuis, O., Gadau, J., Colbourne, J.K.,…Mazur, D.J…..Logsdon, J.M., Jr….Schurko,
A.M….et al. (157 total authors). 2010. Functional and evolutionary insights from the genomes of three parasitoid Nasoniaspecies. Science 343-348.
- Schurko, A.M., Neiman, M. and Logsdon, Jr., J.M. 2009. Signs of sex: what we know and how we know it. Trends in Ecology and Evolution 24:208-217.
- Schurko, A.M., Logsdon, Jr., J.M. and Eads, B.D. 2009. Meiosis genes in Daphnia pulexand the role of parthenogenesis in genome evolution. BMC Evolutionary Biology 9:78.
- Malik S.-B., Pightling A.W., Stefaniak L.M., Schurko
A.M., Logsdon, Jr., J.M. 2008. An expanded inventory of conserved meiotic genes provides evidence for sex in Trichomonas
vaginalis.PLoS One 3:8:e2879.
- Schurko, A.M. and Logsdon, Jr., J.M. 2008. Using a meiosis detection toolkit to investigate ancient asexual “scandals” and the evolution of sex. BioEssays
30:579-589.
- Bedard, J.E.J., Schurko,
A.M., de Cock, A.W.A.M. and Klassen, G.R. 2006. Diversity and evolution of 5S rRNA gene family organization in Pythium. Mycological Research 110:86-95.
- Fernando, W.G.G., Zhang, J.X., Chen, C.Q., Remphrey, W.R., Schurko, A.M. and Klassen, G.R. 2005. Molecular and morphological characteristics of Apiosporina morbosa, the causal agent of black knot in Prunus spp. Canadian Journal of
Plant Pathology 27:364-375.
- Li, Y., Kelly, W. G., Logsdon, Jr., J.M., Schurko, A.M., Harfe, B. D., Hill-Harfe, K. L. and Kahn, R. A. 2004. Functional genomic analysis of the ADP-ribosylation factor family of GTPases: phylogeny among diverse eukaryotes and function in C.
elegans. FASEB Journal 18:1834-1850.
- Iranpour, M., Schurko,
A.M., Klassen, G.R., and Galloway, T.D. 2004. DNA fingerprinting of adult tabanids (Diptera: Tabanidae) and their respective egg masses using PCR-restriction fragment profiling. Canadian Entomologist 136:605-619.
- Schurko, A.M., Mendoza, L., de Cock, A.W.A.M., Bedard, J.E.J., and Klassen, G.R. 2004. Development of a species-specific probe for Pythium insidiosum and the diagnosis of pythiosis. Journal of
Clinical Microbiology 42:2411-2418.
- Schurko, A.M., Mendoza L, Lévesque, C.A., Désaulniers, N.L., de Cock, A.W.A.M., and Klassen G.R. 2003. A molecular phylogeny of Pythium insidiosum. Mycological Research107: 537-544.
- Schurko, A.M.,Mendoza, L., de Cock, A.W.A.M., and Klassen, G.R. 2003. Evidence for geographic clusters: Molecular genetic differences among strains of Pythium insidiosum from Asia, Australia, and the Americas are explored. Mycologia 95:200-208.
Photos of Undergraduate students presenting research at conferences:


